Part:BBa_K3071014
gumB CBS I & II-regulated pspA promoter
Description
This synthetic promoter is developed by fusing the sigma 54-promoter pspA(BBa_K3071013) with CBS I (BBa_K3071011) and CBS II (BBa_K3071012). The activation is based on the interaction of a fusion transction activator pspF TAD-Clp (BBa_K3071009).
Biology
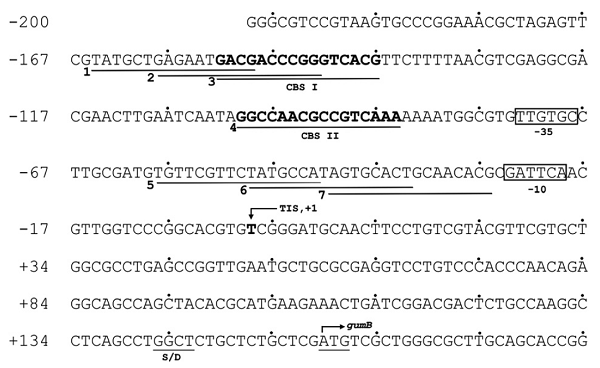
Clp binding sites (CBSs) have typically been identified by pattern searching of the Xcc genome using the consensus CRP binding sequence. Clp upregulates the gum operon by binding to two non-consensus sites. CBS II has a high GC content in the central region (6bp) that may be important for binding, and binding may be enhanced if the GC-rich central region is palindromic.

pspA promoter is a sigma-54 (σ-54) regulated activator dependent promoter. In the orginal pspA promoter upstream region, it contains σ-54 consensus sequence 5' of the start contains a GG doublet as -24 and a consensus GG doublet at -12, the high-affinity IHF site (-25 to -60), as well as the UAS sites (UAS I: -89 to -107; UAS II:-111 to -129).
Sigm54-RNA holoenzyme (σ-54 RNAP) forms an inactive transcriptional initiation complex on this promoter, which can be activated in E. coli by the bacterial enhancer-binding protein PspF (BBa_K3071006). PspF functions by binding to the upstream activation sequences (UAS) near the promoter and contacting the promoter-bound σ-54 RNAP via DNA looping stabilized by the binding of integration host factor (IHF). Previous research has demonstrated the property of PspF-dependent and enhancer-specific transcription activation of pspA promoter.
Usage

The UAS sites of this promoter are replaced by CBS I(BBa_K3071011) and CBSII (BBa_K3071012) in our own synthetic system to construct CBS I & II-regulated pspA promoter (BBa_K3071014). It is the DNA sequence that could interact with pspF-Clp (BBa_K3071009) and lead to the activation of synthetic promoter (BBa_K3071014).

The detail activation of this simga54-dependent promtor is shown in figure 3. sigm54-RNA holoenzyme (σ-54 RNAP) forms an inactive transcriptional initiation complex on this promoter, which can be activated in E. coli by the bacterial enhancer-binding protein PspF (BBa_K3071006). PspF functions by binding to the upstream activation sequences (UAS) near the promoter and contacting the promoter-bound σ-54 RNAP via DNA looping stabilized by the binding of integration host factor (IHF). Previous research has demonstrated the property of pspF-dependent and enhancer-specific transcription activation of pspA promoter.
Characterization
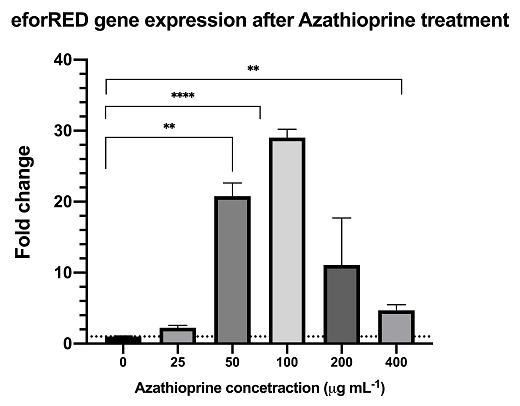
Azathioprine is an indirect supressor to cyclic-di-GMP, treatment of azathioprine to bacteria culture can reduce the cyclic-di-GMP level and lead to activation of the pspF-Clp and by-pass the RpfC/RpfG two-component system.The result from rt-qPCR data shows that the above synthetic biological system is functional with significant up-regulation of reporter mRNA.

The result from rt-qPCR data shows that the above synthetic biological system is functional with significant up-regulation of reporter mRNA upon DSF activation.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 72
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |
